Windows user space issues with installing R packages
Are you a Microsoft Windows R user? Does your Windows username include a space? Like Firstname Lastname. Then you might occassionally run into issues installing packages due to spaces.
Solutions
You could either re-install Windows with a username that has no spaces such as Lastname ^[This is the case in the Bioconductor Windows build machine where the username is biocbuild as you can see here.], but that’s probably not an easy option. Or you can:
- Edit your
TMPandTEMPenvironment variables to a location with no spaces, likeC:\TEMPfollowing instructions like these ones. - Preferably install
Rat a location with no spaces, likeC:\R, instead of the defaultC:\Program Files^[In the Bioconductor Windows build machines there are again no spaces in the path to the R installation and the library where packages are installed.].
Backstory
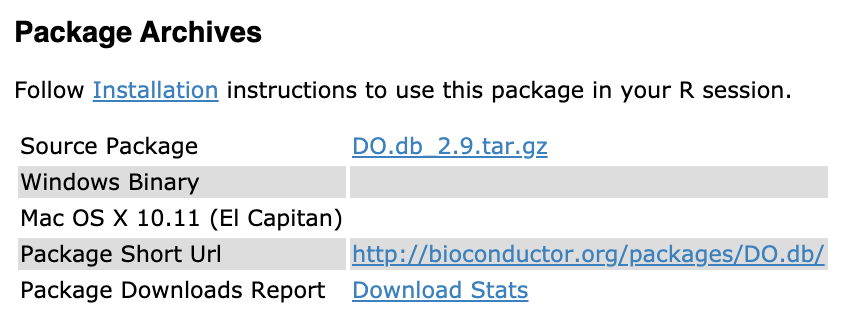
A co-worker wanted to install the clusterprofiler Bioconductor package which depends on the DO.db Bioconductor package. This co-worker uses a Windows machine that has a username with a space. Let’s say it was me with Leo Collado to keep them anonymous. The DO.db is only available as a “Source” package with no Windows binary as you can see here.
{width=600px}
This means that R has to:
- download a
tar.gzfile, - uncompress it,
- and then install it.
In particular, we are talking about DO.db_2.9.tar.gz in this case.
The installation instructions for DO.db are:
if (!requireNamespace("BiocManager", quietly = TRUE))
install.packages("BiocManager")
BiocManager::install("DO.db")
Uncompressing
Internally, BiocManager::install() ends up using utils::install.packages(). The first step, downloading, works well. Uncompressing a file in this scenario fails. Why?
> BiocManager::install('DO.db', lib = 'C:/R/R-3.6.0/library')
Bioconductor version 3.9 (BiocManager 1.30.4), R 3.6.0 (2019-04-26)
Installing package(s) 'BiocVersion', 'DO.db'
## removed output
trying URL 'https://bioconductor.org/packages/3.9/data/annotation/src/contrib/DO.db_2.9.tar.gz'
Content type 'application/x-gzip' length 1769978 bytes (1.7 MB)
downloaded 1.7 MB
Error in untar2(tarfile, files, list, exdir, restore_times) :
incomplete block on file
The downloaded source packages are in
‘C:\Users\Leo Collado\AppData\Local\Temp\RtmpqiBJ53\downloaded_packages’
If you search on Google the error message you’ll find links like this one which hint towards an incomplete download. But the download works. You can even download the file and try to run untar() manually and it will fail.
We were told to try installing R at a location with no spaces, so by this point, R was installed at C:\R\R-3.6.0\, hence the lib specification you see above, though it’s irrelevant for these errors.
Uncompressing the tar.gz file is done by utils::untar(). If you look at the code for utils::untar() you’ll see:
## The function definition of utils::untar
function (tarfile, files = NULL, list = FALSE, exdir = ".", compressed = NA,
extras = NULL, verbose = FALSE, restore_times = TRUE,
support_old_tars = Sys.getenv("R_SUPPORT_OLD_TARS",
FALSE), tar = Sys.getenv("TAR"))
## Inside utils::untar()
if (inherits(tarfile, "connection") || identical(tar, "internal")) {
if (!missing(compressed))
warning("argument 'compressed' is ignored for the internal method")
return(untar2(tarfile, files, list, exdir, restore_times))
}
## Further below
TAR <- tar
if (!nzchar(TAR) && .Platform$OS.type == "windows" && nzchar(Sys.which("tar.exe")))
TAR <- "tar.exe"
if (!nzchar(TAR) || TAR == "internal")
return(untar2(tarfile, files, list, exdir))
In this case, the first untar2() call is called. That is: return(untar2(tarfile, files, list, exdir, restore_times)). The error message incomplete block on file is not really informative in this case because untar2() is not happy when there’s a space in the path to the file ^[Hopefully in the future Google will lead you to this blog post and you might avoid the rabbit hole I went through!].
We can get around this untar2() issue by uncompressing the tar.gz file ourselves in a path that has no spaces. For example, if we download DO.db_2.9.tar.gz to C:\R we can uncompress the tar.gz file with:
utils::untar('C:/R/DO.db_2.9.tar.gz')
Installation
Let’s proceed to installing the package.
> install.packages('C:/R/DO.db', repos = NULL, type = 'source', lib = 'C:/R/R-3.6.0/library')
* installing *source* package 'DO.db' ...
** using staged installation
** R
** inst
** byte-compile and prepare package for lazy loading
ARGUMENT 'Collado\AppData\Local\Temp\Rtmp8EQDjB\Rin2ef05088650f' __ignored__
Error: object 'ÿþ' not found
Execution halted
ERROR: lazy loading failed for package 'DO.db'
* removing 'C:/R/R-3.6.0/library/DO.db'
Warning message:
In install.packages("C:/R/DO.db", repos = NULL, type = "source", :
installation of package ‘C:/R/DO.db’ had non-zero exit status
>
Oh noes! It didn’t work 😖 What happened?
If you look closely, you’ll see that it says ARGUMENT 'Collado\AppData\Local\Temp\Rtmp8EQDjB\Rin2ef05088650f' __ignored__. Wait! Collado is the Lastname portion of the username! So we have another space issue ^[By the way, at this point I thought that the error was related to Error: object 'ÿþ' not found and maybe some encoding issues since the DO.db package has Chinese characters.]. That structure though looks very familiar, it’s from base::tempdir()!
> tempdir()
[1] "C:\\Users\\Leo Collado\\AppData\\Local\\Temp\\RtmpqiBJ53"
The help file for ?tempdir contained the clues to solving this issue.
By default, tmpdir will be the directory given by tempdir(). This will be a subdirectory of the per-session temporary directory found by the following rule when the R session is started. The environment variables TMPDIR, TMP and TEMP are checked in turn and the first found which points to a writable directory is used: if none succeeds ‘/tmp’ is used. The path should not contain spaces. Note that setting any of these environment variables in the R session has no effect on tempdir(): the per-session temporary directory is created before the interpreter is started.
We can set the TMPDIR environment variable which will be used by the R session spawned inside the installation of DO.db and… it works!
> Sys.setenv(TMPDIR = 'C:/R/tmp_leo')
> Sys.getenv('TMPDIR')
[1] "C:/R/tmp_leo"
>
>
> install.packages('C:/R/DO.db', repos = NULL, type = 'source', lib = 'C:/R/R-3.6.0/library')
* installing *source* package 'DO.db' ...
** using staged installation
** R
** inst
** byte-compile and prepare package for lazy loading
** help
*** installing help indices
converting help for package 'DO.db'
finding HTML links ... done
DOANCESTOR html
DOBASE html
DOCHILDREN html
DOMAPCOUNTS html
DOOBSOLETE html
DOOFFSPRING html
DOPARENTS html
DOSYNONYM html
DOTERM html
DOTerms-class html
DOTermsAnnDbBimap html
DO_dbconn html
** building package indices
** testing if installed package can be loaded from temporary location
*** arch - i386
*** arch - x64
** testing if installed package can be loaded from final location
*** arch - i386
*** arch - x64
** testing if installed package keeps a record of temporary installation path
* DONE (DO.db)
Making 'packages.html' ... done
clusterProfiler installation
Now we can continue and install clusterProfiler, right?
> BiocManager::install('clusterProfiler', lib = 'C:/R/R-3.6.0/library')
Bioconductor version 3.9 (BiocManager 1.30.4), R 3.6.0 (2019-04-26)
Installing package(s) 'clusterProfiler'
also installing the dependencies ‘sys’, ‘formatR’, ‘askpass’, ‘farver’, ‘backports’, ‘zeallot’, ‘lambda.r’, ‘futile.options’, ‘curl’, ‘mime’, ‘openssl’, ‘hms’, ‘triebeard’, ‘tweenr’, ‘polyclip’, ‘RcppEigen’, ‘colorspace’, ‘utf8’, ‘vctrs’, ‘futile.logger’, ‘snow’, ‘data.table’, ‘fastmatch’, ‘stringr’, ‘httr’, ‘jsonlite’, ‘progress’, ‘urltools’, ‘xml2’, ‘gridGraphics’, ‘ggforce’, ‘ggrepel’, ‘viridis’, ‘labeling’, ‘munsell’, ‘R6’, ‘cli’, ‘crayon’, ‘fansi’, ‘pillar’, ‘BiocParallel’, ‘fgsea’, ‘reshape2’, ‘cowplot’, ‘europepmc’, ‘ggplotify’, ‘ggraph’, ‘ggridges’, ‘gridExtra’, ‘igraph’, ‘purrr’, ‘RColorBrewer’, ‘UpSetR’, ‘gtable’, ‘lazyeval’, ‘rlang’, ‘scales’, ‘tibble’, ‘viridisLite’, ‘withr’, ‘dplyr’, ‘glue’, ‘stringi’, ‘tidyselect’, ‘DOSE’, ‘enrichplot’, ‘ggplot2’, ‘GO.db’, ‘GOSemSim’, ‘plyr’, ‘qvalue’, ‘rvcheck’, ‘tidyr’
## Delete more output
The downloaded binary packages are in
C:\Users\Leo Collado\AppData\Local\Temp\RtmpqiBJ53\downloaded_packages
installing the source packages ‘pillar’, ‘GO.db’
trying URL 'https://cloud.r-project.org/src/contrib/pillar_1.4.1.tar.gz'
Content type 'application/x-gzip' length 228572 bytes (223 KB)
downloaded 223 KB
trying URL 'https://bioconductor.org/packages/3.9/data/annotation/src/contrib/GO.db_3.8.2.tar.gz'
Content type 'application/x-gzip' length 31820866 bytes (30.3 MB)
downloaded 30.3 MB
Error in untar2(tarfile, files, list, exdir, restore_times) :
incomplete block on file
Error in untar2(tarfile, files, list, exdir, restore_times) :
incomplete block on file
The downloaded source packages are in
‘C:\Users\Leo Collado\AppData\Local\Temp\RtmpqiBJ53\downloaded_packages’
The issue is again that utils:::untar2() and thus utils::untar() does not like spaces in the paths. If we look at where the packages were downloaded more closely, we can see a space there at C:\Users\Leo Collado\AppData\Local\Temp\RtmpqiBJ53\downloaded_packages. If you check the help file for utils::install.packages() you’ll see that destdir controls this:
destdir
directory where downloaded packages are stored. If it is NULL (the default) a subdirectory downloaded_packages of the session temporary directory will be used (and the files will be deleted at the end of the session).
If we dig into utils::install.packages() we can see how this comes to play.
## Part of utils::install.packages()
if (is.null(destdir) && nonlocalrepos) {
tmpd <- file.path(tempdir(), "downloaded_packages")
if (!file.exists(tmpd) && !dir.create(tmpd))
stop(gettextf("unable to create temporary directory %s",
sQuote(tmpd)), domain = NA)
}
Setting the environment variable TMPDIR doesn’t work here as the instructions for tempdir() specify ^[Didn’t stop me from trying hehe. I tried using usethis::edit_r_profile() and adding Sys.setenv(TMPDIR = 'C:/R/tmp_leo') but that didn’t work.] although I now see that you can edit the .Renviron file as instructed here.
In any case, if we specify a destdir without spaces we overide the need to control tempdir(), enable utils::untar() to work and we can finally install clusterProfiler 🎉.
> BiocManager::install('clusterProfiler', lib = 'C:/R/R-3.6.0/library', destdir = 'C:/R/dest_leo')
Bioconductor version 3.9 (BiocManager 1.30.4), R 3.6.0 (2019-04-26)
Installing package(s) 'clusterProfiler'
also installing the dependency ‘GO.db’
trying URL 'https://bioconductor.org/packages/3.9/bioc/bin/windows/contrib/3.6/clusterProfiler_3.12.0.zip'
Content type 'application/zip' length 623524 bytes (608 KB)
downloaded 608 KB
package ‘clusterProfiler’ successfully unpacked and MD5 sums checked
installing the source package ‘GO.db’
trying URL 'https://bioconductor.org/packages/3.9/data/annotation/src/contrib/GO.db_3.8.2.tar.gz'
Content type 'application/x-gzip' length 31820866 bytes (30.3 MB)
downloaded 30.3 MB
* installing *source* package 'GO.db' ...
** using staged installation
** R
** inst
** byte-compile and prepare package for lazy loading
** help
*** installing help indices
converting help for package 'GO.db'
finding HTML links ... done
GOBASE html
GOBPANCESTOR html
GOBPCHILDREN html
GOBPOFFSPRING html
GOBPPARENTS html
GOCCANCESTOR html
GOCCCHILDREN html
GOCCOFFSPRING html
GOCCPARENTS html
GOMAPCOUNTS html
GOMFANCESTOR html
GOMFCHILDREN html
GOMFOFFSPRING html
GOMFPARENTS html
GOOBSOLETE html
GOSYNONYM html
GOTERM html
GO_dbconn html
** building package indices
** testing if installed package can be loaded from temporary location
*** arch - i386
*** arch - x64
** testing if installed package can be loaded from final location
*** arch - i386
*** arch - x64
** testing if installed package keeps a record of temporary installation path
* DONE (GO.db)
Making 'packages.html' ... done
Closing
All of the above seemed like too much. In addition, it seemed like BiocManager::install('hypeR', destdir = 'C:/R/dest_leo') was not working ^[I would need to test this more before reporting it properly to Bioconductor.]. I likely missed something here earlier today. So controlling utils::tempdir() seemed like the easiest solution such that the defaults of where a package gets downloaded, uncompressed, etc all worked. And the simplest solution we thought of was to create the C:\TEMP directory and update the Windows environment variables TMP and TEMP to point to that location. Then, the rest of the commands worked without having to specify lib or destdir or manually run utils::untar().
As a whole, remember to look for spaces in the error messages! This is specially relevant when you are having issues as a Microsoft Windows R user.
If you have other solutions for Microsoft Windows R users with usernames that have at least one space, please let us know in the comments! Thank you! 🙌🏽
Acknowledgments
This blog post was made possible thanks to:
- BiocStyle (Oleś, 2023)
- blogdown (Xie, Hill, and Thomas, 2017)
- knitcitations (Boettiger, 2021)
- sessioninfo (Wickham, Chang, Flight, Müller et al., 2021)
References
[1] C. Boettiger. knitcitations: Citations for 'Knitr' Markdown Files. R package version 1.0.12. 2021. URL: https://CRAN.R-project.org/package=knitcitations.
[2] A. Oleś. BiocStyle: Standard styles for vignettes and other Bioconductor documents. R package version 2.28.0. 2023. DOI: 10.18129/B9.bioc.BiocStyle. URL: https://bioconductor.org/packages/BiocStyle.
[3] H. Wickham, W. Chang, R. Flight, K. Müller, et al. sessioninfo: R Session Information. R package version 1.2.2. 2021. URL: https://CRAN.R-project.org/package=sessioninfo.
[4] Y. Xie, A. P. Hill, and A. Thomas. blogdown: Creating Websites with R Markdown. Boca Raton, Florida: Chapman and Hall/CRC, 2017. ISBN: 978-0815363729. URL: https://bookdown.org/yihui/blogdown/.
Reproducibility
## ─ Session info ───────────────────────────────────────────────────────────────────────────────────────────────────────
## setting value
## version R version 4.3.1 (2023-06-16)
## os macOS Ventura 13.4
## system aarch64, darwin20
## ui X11
## language (EN)
## collate en_US.UTF-8
## ctype en_US.UTF-8
## tz America/New_York
## date 2023-07-11
## pandoc 3.1.5 @ /opt/homebrew/bin/ (via rmarkdown)
##
## ─ Packages ───────────────────────────────────────────────────────────────────────────────────────────────────────────
## package * version date (UTC) lib source
## backports 1.4.1 2021-12-13 [1] CRAN (R 4.3.0)
## bibtex 0.5.1 2023-01-26 [1] CRAN (R 4.3.0)
## BiocManager 1.30.21 2023-06-10 [1] CRAN (R 4.3.0)
## BiocStyle * 2.28.0 2023-04-25 [1] Bioconductor
## blogdown 1.18 2023-06-19 [1] CRAN (R 4.3.0)
## bookdown 0.34 2023-05-09 [1] CRAN (R 4.3.0)
## bslib 0.5.0 2023-06-09 [1] CRAN (R 4.3.0)
## cachem 1.0.8 2023-05-01 [1] CRAN (R 4.3.0)
## cli 3.6.1 2023-03-23 [1] CRAN (R 4.3.0)
## colorout 1.2-2 2023-05-06 [1] Github (jalvesaq/colorout@79931fd)
## digest 0.6.31 2022-12-11 [1] CRAN (R 4.3.0)
## evaluate 0.21 2023-05-05 [1] CRAN (R 4.3.0)
## fastmap 1.1.1 2023-02-24 [1] CRAN (R 4.3.0)
## generics 0.1.3 2022-07-05 [1] CRAN (R 4.3.0)
## glue 1.6.2 2022-02-24 [1] CRAN (R 4.3.0)
## htmltools 0.5.5 2023-03-23 [1] CRAN (R 4.3.0)
## httr 1.4.6 2023-05-08 [1] CRAN (R 4.3.0)
## jquerylib 0.1.4 2021-04-26 [1] CRAN (R 4.3.0)
## jsonlite 1.8.7 2023-06-29 [1] CRAN (R 4.3.0)
## knitcitations * 1.0.12 2021-01-10 [1] CRAN (R 4.3.0)
## knitr 1.43 2023-05-25 [1] CRAN (R 4.3.0)
## lifecycle 1.0.3 2022-10-07 [1] CRAN (R 4.3.0)
## lubridate 1.9.2 2023-02-10 [1] CRAN (R 4.3.0)
## magrittr 2.0.3 2022-03-30 [1] CRAN (R 4.3.0)
## plyr 1.8.8 2022-11-11 [1] CRAN (R 4.3.0)
## R6 2.5.1 2021-08-19 [1] CRAN (R 4.3.0)
## Rcpp 1.0.10 2023-01-22 [1] CRAN (R 4.3.0)
## RefManageR 1.4.0 2022-09-30 [1] CRAN (R 4.3.0)
## rlang 1.1.1 2023-04-28 [1] CRAN (R 4.3.0)
## rmarkdown 2.23 2023-07-01 [1] CRAN (R 4.3.0)
## rstudioapi 0.14 2022-08-22 [1] CRAN (R 4.3.0)
## sass 0.4.6.9000 2023-05-06 [1] Github (rstudio/sass@f248fe5)
## sessioninfo * 1.2.2 2021-12-06 [1] CRAN (R 4.3.0)
## stringi 1.7.12 2023-01-11 [1] CRAN (R 4.3.0)
## stringr 1.5.0 2022-12-02 [1] CRAN (R 4.3.0)
## timechange 0.2.0 2023-01-11 [1] CRAN (R 4.3.0)
## xfun 0.39 2023-04-20 [1] CRAN (R 4.3.0)
## xml2 1.3.4 2023-04-27 [1] CRAN (R 4.3.0)
## yaml 2.3.7 2023-01-23 [1] CRAN (R 4.3.0)
##
## [1] /Library/Frameworks/R.framework/Versions/4.3-arm64/Resources/library
##
## ──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────