Teaching a short topic to beginner R users
A couple weeks ago I was given the opportunity to teach a 1 hr 30 min slot of an introduction to R course. In the past, I’ve taught lectures for similar courses, and I ended up asking myself what would be the best short topic to teach and how to teach it.
Best short topic
There are two ways to answer the first question, one boring and one more interesting. The boring answer is that the course instructor selected the topic. The interesting one goes like this. I have taken short R courses before and taught others, and it’s always overwhelming for the students. You get to cover many concepts, get familiarized with R’s syntax, and in the end without lots of practice it’s very challenging to retain much of the information. I think that students love it when they learn how to do something simple that could be the first building block for many of their projects. In parallel, I think that one of the coolest R topics you can learn in an hour is how to create reproducible documents with rmarkdown (Allaire, Cheng, Xie, McPherson, et al., 2015).
Learning how to use a single function, render() in this case, is as simple as it gets. And using the RStudio Desktop is even simpler. Of course, it can easily get complicated. For example, on a new computer you need to install all the LaTeX dependencies if you want to create PDF files. That task can take some time and maybe scare away some new users. But PDF files are really a plus in this case since you can start creating HTML and Word documents. Other complications arise when a user is interested in more control over formatting the file, but like I said earlier, all you need is a simple building block and rmarkdown is clearly one of them.
This is why the final answer to the first question was teaching how to use rmarkdown to create reproducible reports (HTML, Word files) using R.
How to teach it
Teaching a short topic to a beginner’s audience is no easy feat. In the past I’ve made lectures that have the code for every single step and many links to resources where students can learn some details. That is, I’ve created the lectures in such a way that a student can later use them as reference and follow them without an instructor explaining them.
That’s a strategy that I think works on the long run. However, it makes the actual lecture boring and very limited in interactivity. At the JHSPH biostat computing club, other students have chosen to use a lot of images, funny to witty quotes, and asked listeners to voice their opinions. I’ve come to enjoy those presentations and I decided to create my lecture following that trend.
I started off with a series of questions about reproducible research and asked students to voice their opinions and to define a few key concepts. A couple were aware of the difference between reproducibility and replicability, but most were not. I also questioned them and presented them verbally with some famous cases, so they could realize that it’s a fairly complicated matter. Next I presented some answers and definitions from the Implementing Reproducible Research book.
Specifically talking about R, I showed the students several documents I’ve created in the past and asked whether they thought that they could reproduce the results or not. Basically, I wanted to highlight that when using R, you really need the session information if you want to reproduce something. Specially if the analysis involves packages under heavy development.
After having motivating the need for reproducible documents, I briefly showed what rmarkdown is with some images from RStudio shown below.
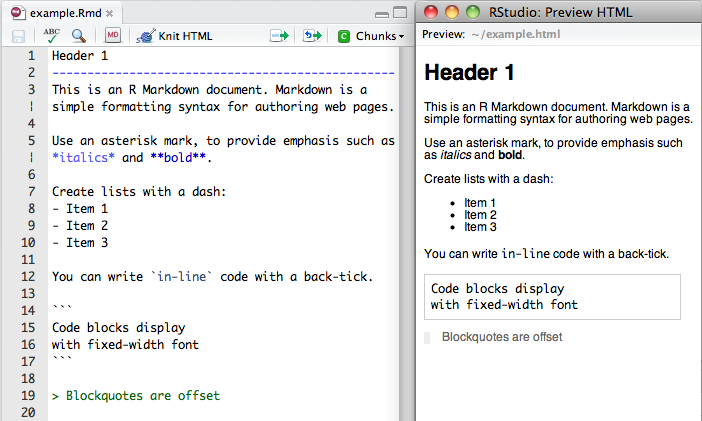
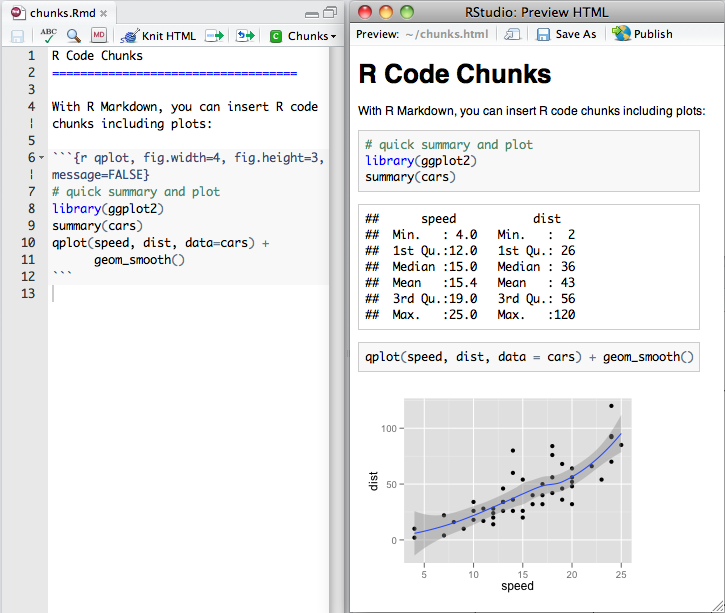
That gave the students a general idea of how these documents look when you are writing them. But the most important part was showing them examples of how the resulting documents look like. That is, I showed them some complicated projects so they could imagine doing one themselves. The examples included some books, but given the audience I think that the one that motivated them most was Alyssa Frazee’s polyester reproducible paper (check the source here). I also showed them some of the cool stuff you can create with HTML documents: basically adding interactive elements.
From there, we left the presentation and I demo’ed how to use RStudio to write rmarkdown documents, the Markdown syntax, where to find help, etc.

By this point, I think the lecture was quite complete and the students were motivated. However, from my past experience, I’ve come to realize that students will easily forget a topic if they don’t practice doing it. That is why even before making the lecture I spent quite a bit of time designing two practice labs. Both labs involved creating a rmarkdown document.
The first lab included some cool illusion plots which involved a lot of R code. The code wasn’t the point, but simply learning some of the basics such as what is a code chunk, some of Markdown’s syntax, specifying some code chunk options, adding the session information, and using inline R code to show the date when the document was made. Ahh, and of course, uploading your HTML document to RPubs (see mine). I know that not everyone is a fan of RPubs, but I imagined that students would get super excited that they made something that they could then show their colleagues and friends. And some did!
Sadly, we didn’t have enough time for the second lab. I did explain to the students what it was about, but they didn’t have time to do it themselves. For this second document, I wanted the students to learn how to create a document reporting some results where all the numbers in the text are written by R instead of copy-pasting them.
Conclusions
As you can see, I enjoyed thinking what to teach and specially how to teach a short topic to beginner R students. Thanks to having one of the later sessions, I could teach them how to use rmarkdown in a way that hopefully left them highly motivated to do it themselves. I hope that most of them will take that they learned in that module and others and apply them in their day to day work.
References
You can find the lecture itself here but like I said earlier, it was designed for class and not for being used as a reference. However, the lab and it’s key might be more useful.
Citations made with knitcitations (Boettiger, 2015).
[1] J. Allaire, J. Cheng, Y. Xie, J. McPherson, et al. rmarkdown: Dynamic Documents for R. R package version 0.7. 2015. URL: http://CRAN.R-project.org/package=rmarkdown.
[2] C. Boettiger. knitcitations: Citations for Knitr Markdown Files. R package version 1.0.6. 2015. URL: http://CRAN.R-project.org/package=knitcitations.
Want more?
Check other @jhubiostat student blogs at Bmore Biostats as well as topics on #rstats.