Translational Neuroscience Division, Data Science I
JHPCE: lieber_lcolladotor
Lieber Institute for Brain Development
R/Bioconductor-powered Team Data Science

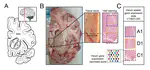
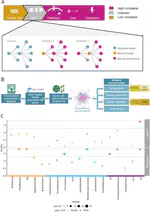
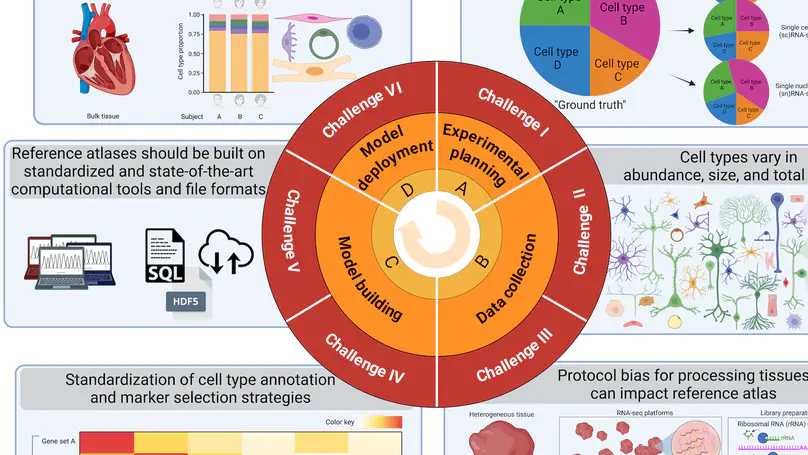
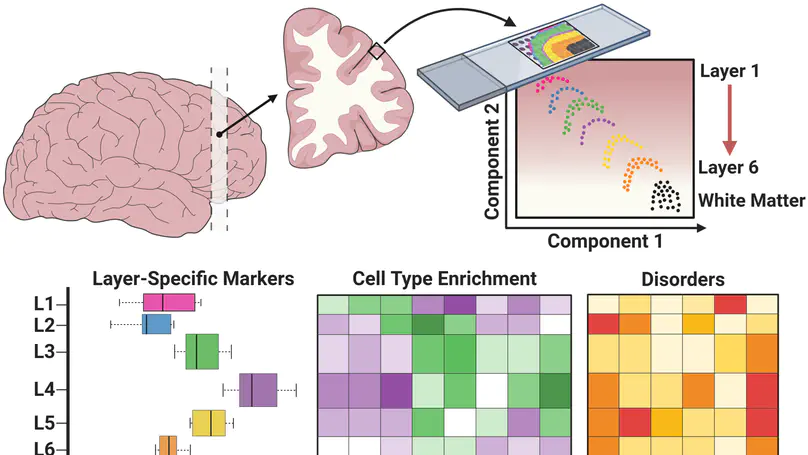
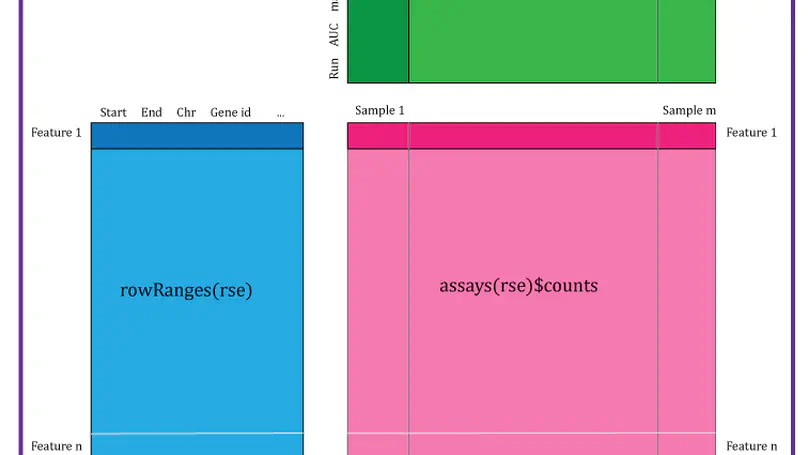
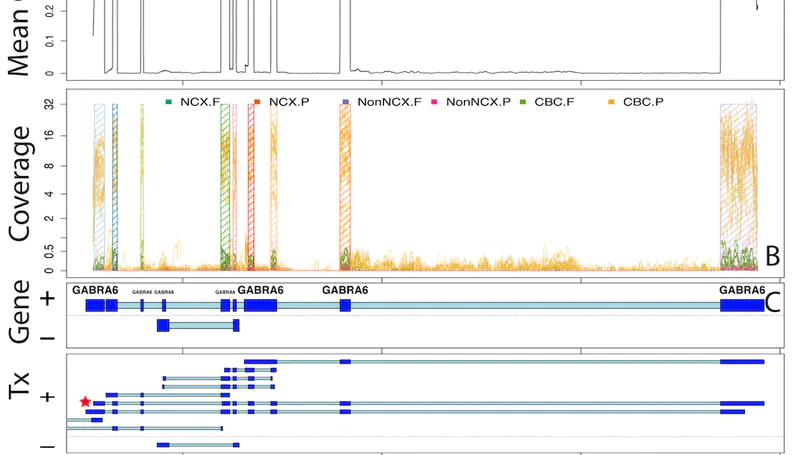
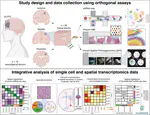
At the Lieber Institute for Brain Development (LIBD), as part of the Translational Neuroscience Division, our group works on understanding the roots and signatures of disease (particularly psychiatric disorders) by zooming in across dimensions of gene activity. We achieve this by studying gene expression at all expression feature levels (genes, exons, exon-exon junctions, and un-annotated regions) and by using different gene expression measurement technologies (bulk RNA-seq, single cell/nucleus RNA-seq, and spatial transcriptomics) that provide finer biological resolution and localization of gene expression. We work closely with collaborators from LIBD as well as from Johns Hopkins University (JHU), University of Cambridge, and other institutions, which reflects the cross-disciplinary approach and diversity in expertise needed to further advance our understanding of high throughput biology.
In order to provide a supportive and stimulating research environment at LIBD, our group provides Data Science guidance sessions open to any LIBD staff member and we organize the LIBD rstats club, among other initiatives. Our documentation book website contains more details for on boarding, how to ask for help, bootcamps, writing papers, authorship, configuration files, and much more.
Check out the content we share
We constantly create new content to share what we are learning or working on, which you might be interested in. In particular, we:
- run the LIBD rstats club: rstats club schedule
- discuss papers and software in our team meetings: team meeting schedule
Join the team
If you are interested in joining the R/Bioconductor-powered Team Data Science group, please check our open positions at the LIBD career opportunities website. You might be interested in checking our anonymous team survey results, which highlights some strengths but also some weaknesses and areas we can improve.
If we don’t have any open positions, please reach out to Leonardo with your CV, GitHub/GitLab/etc profile with open-source software, and a short description of why you are interested in our team.
Fields
that drive us
Projects
All talks
Leonardo’s recent publications
and posters
† indicates corresponding author, * indicates equal contribution
You can also find Leonardo’s publications list at NCBI, ORCiD, and Google Scholar.
Leonardo’s recent blog posts
Posts with the rstats category can also be found at RBloggers and R Weekly. Also check the LIBD rstats club where Leonardo is a contributor. You can also view posts grouped by category or tag.
Courses taught by Leonardo
LIBD

2024
- Instructor of the Introduction to RNA-seq data analysis with Bioconductor course for LCG-UNAM students.
- Invited instructor for this EMBL-EBI virtual course.
- Instructor for a portion of the Statistical Analysis of Genome Scale Data course at Cold Spring Harbor Laboratory.
- Instructor for 140.776 Statistical Computing course at the Johns Hopkins Bloomberg School of Public Health.
2023
- Instructor of the Introduction to RNA-seq data analysis with Bioconductor course for LCG-UNAM students.
- Instructor for a portion of the Statistical Analysis of Genome Scale Data course at Cold Spring Harbor Laboratory.
- Instructor for 140.776 Statistical Computing course at the Johns Hopkins Bloomberg School of Public Health.
- Invited instructor for CDSB 2023.
- Invited instructor for this PROINNOVA-funded course for Centro Universitario Los Altos.
2022
- Instructor of the Introduction to RNA-seq data analysis with Bioconductor course for LCG-UNAM students.
2021
- Organizer and instructor for the CDSB 2021 workshop: analysis of scRNA-seq data with Bioconductor.
- Instructor of the Introduction to RNA-seq data analysis with Bioconductor course for LCG-UNAM students.
- Instructor of the Getting started with scRNA-seq analyses with Bioconductor 2 hour workshop for the Human Cell Atlas - Latin America workshop.
- Instructor of the Interactive exploration of RNA-seq data with iSEE mini course that is part of the mini courses series organized by CDSB, RMB and NNB-CCG (UNAM).
2020
- Instructor for the R/Bioconductor Data Science LIBD bootcamps.
- Instructor and member of the Organizing Committee for the CDSB Workshop 2020: Building workflows with RStudio and Bioconductor for single cell RNA-seq analysis

- Instructor of the Analyzing scRNA-seq data with Bioconductor for LCG-EJ-UNAM students.
2019
- Instructor and member of the Organizing Committee for the CDSB Workshop 2019: How to Build and Create Tidy Tools

2018
- Keynote speaker and member of the Organizing Committee for the Latin American R/BioConductor Developers Workshop 2018

2016
- Biostatistics and Stata instructor at a workshop for Kandahar University Faculty, organized by Johns Hopkins University.
- Invited instructor for the Genomeeting 2016 course taught at INMEGEN, Mexico City, Mexico.
JHBSPH

2015-2016
- Teaching assistant and guest lecturer for Introduction to R for Public Health Researchers.
- Teaching assistant for Statistical Methods in Public Health I (140.621).
- Lead teaching assistant for Statistical Methods in Public Health II (140.622).
- Teaching assistant for the MPH capstone project.
2014-2015
- Lead teaching assistant for Statistical Methods in Public Health I (140.621) and II (140.622).
- Teaching assistant for the MPH capstone project.
2013-2014
- Teaching assistant for Statistical Methods in Public Health I (140.621) and II (140.622).
- Teaching assistant for the MPH capstone project. Developed a shiny application that allows students to sign up for a TA session (code) and wrote a report of the number of TA sessions available here.
2012-2013
- Teaching assistant for Statistical Methods in Public Health I (140.621), II (140.622), III (140.623), and IV (140.624) courses.
UNAM

PDCB
While working at Winter Genomics I taught two courses for students of the Biomedical Sciences PhD Program (PDCB) from the National Autonomous University of Mexico (UNAM).
- Analysis of High-Throughput Sequencing data with Bioconductor Aug-Dec 2010.
- Introduction to R and Biostatistics (along with two other teachers).
IBT
While I was at the Institute of Biotechnology (UNAM) working with the Winter Genomics crew I organized two courses. One was a series of various bioinformatics and biology mini-courses and another one involved members of different academic institutions.
- Introduction to R for bench biologists Oct-Nov 2009. This mini-course has quite a bit of material on learning how to make plots with R.
- Statistical Methods and Analysis of Genomic Data Jan 2010. This one week course had lectures about Perl, using a Cluster, high-throughput technologies, R and Bioconductor, C, and biology overviews.
LCG
I taught three courses during my undergrad stage at the Undergraduate Program on Genomic Sciences (LCG). Each of these courses has its own website organizing the material. These are:
- Intensive course on R/Bioconductor Oct-Nov 2008
- Principles of Statistics Feb-June 2009
- Seminar III: R/Bioconductor Aug-Dec 2009
Leonardo’s curriculum vitae
Download Leonardo’s cv or view it on GitHub.
Contact
If you have questions about the R/Bioconductor packages that Leonardo maintains, please read this post. If you send him an email, he’ll simply refer you to the same blog post.
- lcolladotor@gmail.com
- +1-301-450-2083
- 855 N. Wolfe, Room 382, Baltimore, MD 21205
- Enter the Rangos building, register at the Security fron desk, take the elevator to the third floor, and register at the LIBD front desk.
- Book an appointment
- DM Leonardo