spatial
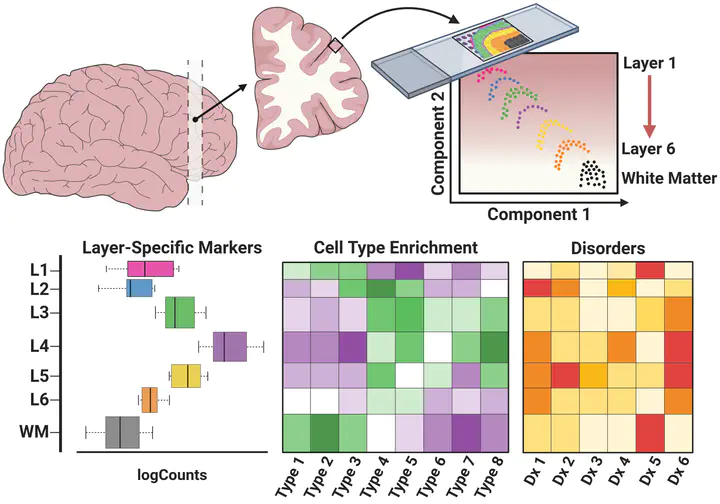 Image credit: bioRxiv
Image credit: bioRxiv
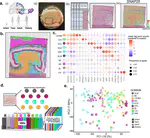
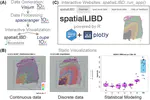
In February 2020 we published the first pre-print using the 10x Genomics Visium platform for spatial transcriptomics. For this project we created the spatialLIBD Bioconductor package and started developing new analytical methods for this type of data. Given that we shared all the data and code when we posted the pre-print, by the time the peer-reviewed publication was made public in February 2021, others had already used our dataset to develop and showcase their methods. For example, BayesSpace’s pre-print was posted on September 2020, prior to their own June 2021 peer-reviewed publication. We are now using BayesSpace in our projects. This is an example of how open science and open access accelerate science.
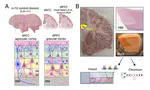
🔥off the press! 👀 our @biorxivpreprint on human 🧠brain @LieberInstitute spatial 🌌🔬transcriptomics data 🧬using Visium @10xGenomics🎉#spatialLIBD
— 🇲🇽 Leonardo Collado-Torres (@lcolladotor) February 29, 2020
🔍https://t.co/RTW0VscUKR
👩🏾💻https://t.co/bsg04XKONr
📚https://t.co/FJDOOzrAJ6
📦https://t.co/Au5jwADGhYhttps://t.co/PiWEDN9q2N pic.twitter.com/aWy0yLlR50
Our paper describing our package #spatialLIBD is finally out! 🎉🎉🎉
— Brenda Pardo (@PardoBree) April 30, 2021
spatialLIBD is an #rstats / @Bioconductor package to visualize spatial transcriptomics data.
⁰
This is especially exciting for me as it is my first paper as a first author 🦑.https://t.co/COW013x4GA
1/9 pic.twitter.com/xevIUg3IsA
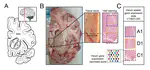
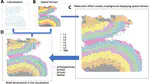
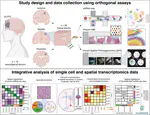
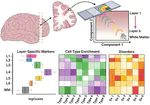
Hot of the pre-print press! 🔥 Our latest work #spatialDLPFC pairs #snRNAseq and #Visium spatial transcriptomic data in the human #DLPFC building a neuroanatomical atlas of this critical brain region 🧠@LieberInstitute @10xGenomics #scitwitter
— Louise Huuki-Myers (@lahuuki) February 17, 2023
📰 https://t.co/NJWJ1mwB9J pic.twitter.com/l8W154XZ50
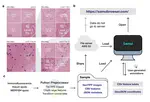
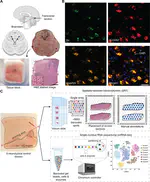
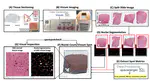
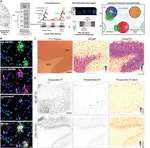
After 10+ years of research, my FIRST 1st-author paper & @biorxivpreprint is finally OUT🥹I delved deep into the human inferior temporal cortex🧠in #Alzheimersdisease, using #VisiumSPG #VectraPolaris #RNAscope & #HALO @Indica_Labs. Check out our📜at https://t.co/QQEy1ZWc1V🔥 pic.twitter.com/dBZNzL1S96
— Sang Ho (Sangho) Kwon (@sanghokwon17) April 24, 2023
Leonardo Collado-Torres
Investigator @ LIBD, Assistant Professor, Department of Biostatistics @ JHBSPH
#rstats @Bioconductor/🧠 genomics @LieberInstitute/@lcgunam @jhubiostat @jtleek @andrewejaffe alumni/@LIBDrstats @CDSBMexico co-founder